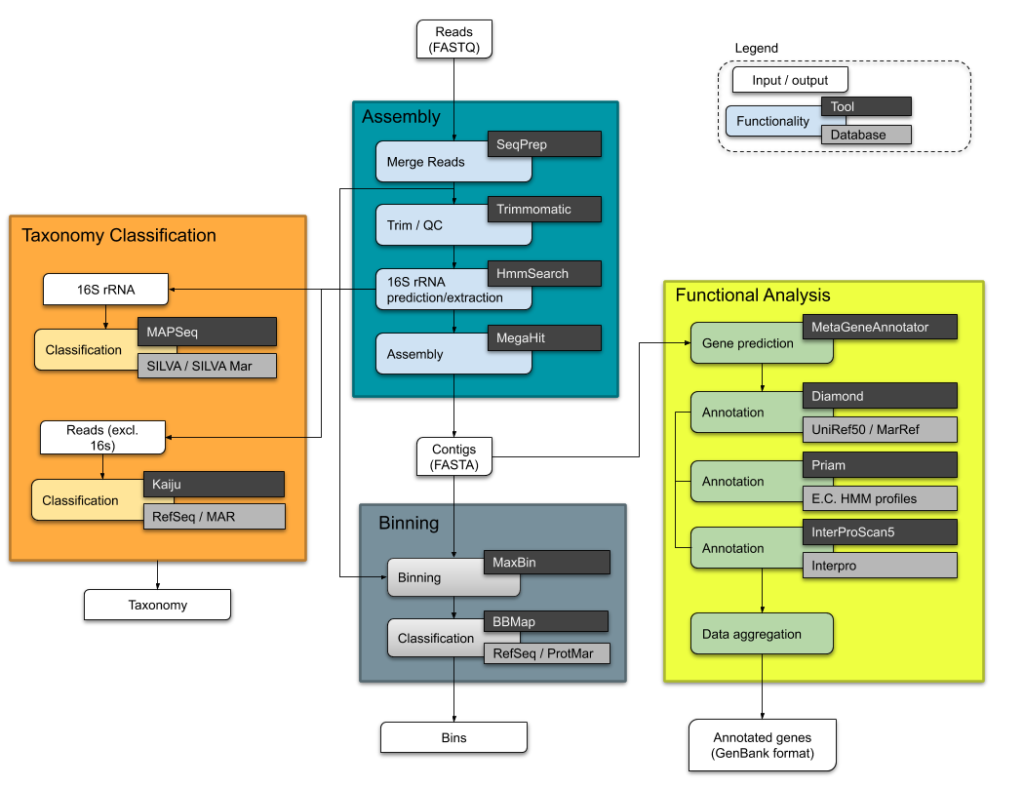
META-pipe is a Nextflow workflow for annotation and analysis of marine metagenomics samples, which provides insight into phylogenetic diversity, metabolic and functional potential of environmental communities. META-pipe consists of four parts; processing of reads (merging, filtering, 16S rRNA extraction and assembly), taxonomic classification (using reads against RefSeq and MAR databases, and predicted 16S rRNA), binning and functional assignment of predicted coding sequences (CDSs) as shown in the figure below.
META-pipe service
META-pipe available as a web service provided by Elixir Norway. Simply upload your metagenomes and we take care of the rest.


